- Blog
- Home
- Best Free Accounting Software For Mac 2017
- Best Mac Document Scanner Software
- Theatre Script Writing Software Mac
- Where Is The Keychain App On A Mac
- Does Wd My Passport Software Work On Mac
- Roland Gx-24 Cutstudio Software Download Mac
- Best Internet Radio Software Mac
- Dji Inspire App For Mac
- Best 3d Print Software Mac
- Tv App Mac Os X
- Gene Marker Software For Mac
- Samsung S8 Unlock Mac Software
- Canon Mg6400 Scanner Software For Mac
- Remote For Mac Mini App
- Electronic Schematic Software Free Mac
- Add Calendat To Mac Today App
- Installing Software Mac Across All Accoumts
- Mac Os X Chromcast App 10.7.5
- Remove The Dymo App On My Mac
- Viper Mouse V560 Software Download Mac
- Free App To Transfer Files From Note 8 To Mac
- Mac App Check Disk Space
- Online Business Tax Software For Mac
- Comic Drawing Software Free Mac
- Mac Os 3d Printing Software
- Adobe After Effects Render Engine App Mac
- House Floor Plan App For Mac
- Tata Photon Software For Mac
- Spectrum App For Mac Pro
- Digital Mixer Software For Mac
- Free Relational Database Software For Mac
- Slideshow Software For Mac 2014
- Why Does Mac Book App Ruin The Book Format
- Free Smart Notebook Software For Mac
- Mac Place App On Desktop
- Photo Manipulation Software For Mac Free
- Texting Apps For Droid And Mac
- Electrical Schematic Software Free Mac
- When Apps Are Downloaded On Macs Where Do They Go
- Force Empty Trash Mac App
- Smart Board Software Download Mac
- Wps Office Presentation Software Mac
- Best Online 2d Animation Software For Mac
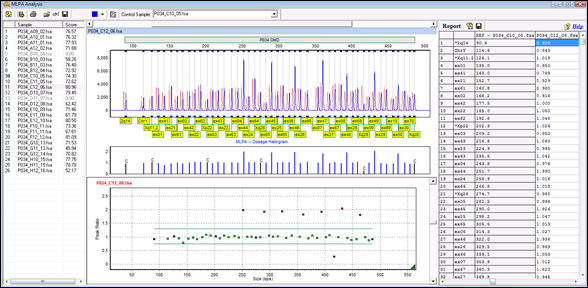
To deal with the increase in information and sensitivity gained from a 24-plex STR amplification kit, the Richland County Sheriff’s Department’s DNA laboratory evaluated and validated a new data analysis software, GeneMarker® HID v 2.7.1. GeneMarker 2 2 + Crack Keygen/Serial Date added: January 2020. Copy Download Link (paste this to your browser).
Chicks atop a picture of a genetic map of a chicken. The chicken genome has 39 pairs of chromosomes, whereas the human genome contains 23 pairs
A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. It can be described as a variation (which may arise due to mutation or alteration in the genomic loci) that can be observed. A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change (single nucleotide polymorphism, SNP), or a long one, like minisatellites.
Background[edit]
For many years, gene mapping was limited to identifying organisms by traditional phenotype markers. This included genes that encoded easily observable characteristics such as blood types or seed shapes. The insufficient number of these types of characteristics in several organisms limited the mapping efforts that could be done. This prompted the development of gene markers which could identify genetic characteristics that are not readily observable in organisms (such as protein variation).[1]
Types[edit]
SFP discovery principle for gene probing
Some commonly used types of genetic markers are:
- RFLP (or Restriction fragment length polymorphism)
- SSLP (or Simple sequence length polymorphism)
- AFLP (or Amplified fragment length polymorphism)
- RAPD (or Random amplification of polymorphic DNA)
- VNTR (or Variable number tandem repeat)
- SSR Microsatellite polymorphism, (or Simple sequence repeat)
- SNP (or Single nucleotide polymorphism)
- STR (or Short tandem repeat)
- SFP (or Single feature polymorphism)
- DArT (or Diversity Arrays Technology)
- RAD markers (or Restriction site associated DNA markers)
Molecular genetic markers can be divided into two classes a) biochemical markers which detect variation at the gene product level such as changes in proteins and amino acids and b) molecular markers which detect variation at the DNA level such as nucleotide changes: deletion, duplication, inversion and/or insertion. Markers can exhibit two modes of inheritance, i.e. dominant/recessive or co-dominant. If the genetic pattern of homo-zygotes can be distinguished from that of hetero-zygotes, then a marker is said to be co-dominant. Generally co-dominant markers are more informative than the dominant markers.[2]
Uses[edit]
Genetic markers can be used to study the relationship between an inherited disease and its genetic cause (for example, a particular mutation of a gene that results in a defective protein). It is known that pieces of DNA that lie near each other on a chromosome tend to be inherited together. This property enables the use of a marker, which can then be used to determine the precise inheritance pattern of the gene that has not yet been exactly localized.
Genetic markers are employed in genealogical DNA testing for genetic genealogy to determine genetic distance between individuals or populations. Uniparental markers (on mitochondrial or Y chromosomal DNA) are studied for assessing maternal or paternal lineages. Autosomal markers are used for all ancestry.
Genetic markers have to be easily identifiable, associated with a specific locus, and highly polymorphic, because homozygotes do not provide any information. Detection of the marker can be direct by RNA sequencing, or indirect using allozymes.
Some of the methods used to study the genome or phylogenetics are RFLP, AFLP, RAPD, SSR. They can be used to create genetic maps of whatever organism is being studied.
There was a debate over what the transmissible agent of CTVT (canine transmissible venereal tumor) was. Many researchers hypothesized that virus like particles were responsible for transforming the cell, while others thought that the cell itself was able to infect other canines as an allograft. With the aid of genetic markers, researchers were able to provide conclusive evidence that the cancerous tumor cell evolved into a transmissible parasite. Furthermore, molecular genetic markers were used to resolve the issue of natural transmission, the breed of origin (phylogenetics), and the age of the canine tumor.[3]
Genetic markers have also been used to measure the genomic response to selection in livestock. Natural and artificial selection leads to a change in the genetic makeup of the cell. The presence of different alleles due to a distorted segregation at the genetic markers is indicative of the difference between selected and non-selected livestock.[4]
References[edit]
- ^Benjamin A. Pierce (2013-12-27). Genetics: A Conceptual Approach. Macmillan Learning. ISBN978-1-4641-0946-1.
- ^N Manikanda Boopathi (2012-12-12). Genetic Mapping and Marker Assisted Selection: Basics, Practice and Benefits. Springer Science & Business Media. pp. 60–. ISBN978-81-322-0958-4.
- ^Murgia C, Pritchard JK, Kim SY, Fassati A, Weiss RA. Clonal origin and evolution of a transmissible cancer. Cell. 2006 Aug 11;126(3):477-87.
- ^Gomez-Raya L, Olsen HG, Lingaas F, Klungland H, Våge DI, Olsaker I, Talle SB, Aasland M, Lien S (November 2002). 'The use of genetic markers to measure genomic response to selection in livestock'. Genetics. 162 (3): 1381–8. PMC1462338. PMID12454081.
See also[edit]
Further reading[edit]
- de Vicente C, Fulton T (2003). Molecular Marker Learning Modules – Vol. 1. IPGRI, Rome, Italy and Institute for Genetic Diversity, Ithaca, New York, USA.[permanent dead link]
- de Vicente C, Fulton T (2004). Molecular Marker Learning Modules – Vol. 2. IPGRI, Rome, Italy and Institute for Genetic Diversity, Ithaca, New York, USA.
- de Vicente C, Glaszmann JC (2006). Molecular Markers for Allele Mining. AMS (Bioversity's Regional Office for the Americas), CIRAD, GCP, IPGRI, M.S. Swaminathan Research Foundation. p. 85. Archived from the original on 2007-12-04. Retrieved 2007-12-12.
- Spooner D, van Treuren R, de Vicente MC (2005). Molecular markers for genebank management. CGN, IPGRI, USDA. p. 126. Archived from the original on 2008-05-03. Retrieved 2007-12-12.
Software For Mac Free
External links[edit]
Media related to Genetic markers at Wikimedia Commons
Retrieved from 'https://en.wikipedia.org/w/index.php?title=Genetic_marker&oldid=949654676'
- Wormbase Converter is a simple, easy to use application designed to help you with the conversion of any Gene ID into any other Gene ID.You can convert a 'WormBase ID', a 'Gene/Transcript/CDS Sequence Name' or a 'Gene Name' into a 'WormBase ID', a. ...
- WBConverter__Local__Windows.zip
- Team J. Ewbank
- Freeware (Free)
- WindowsAll
- The basic allelic case-only test for interaction between a pair of genetic loci. This implementation uses a permutation test to calculate marker-pair-wise p-values, gene-pair-wise p-values, and global family-wise p-values. See the read.me, or. ...
- coip.tar.gz
- coiptestv1
- Freeware (Free)
- 26 Kb
- Windows; Mac; Linux
- Gene Visualizer is a small app designed to display an organism's genes visually. The program takes two sets of genes as input. One is a list of an organism's predicted genes, and the other is a list of an organism's known genes. It can be useful to. ...
- GeneVisualizer.jar
- Andy Hou
- Freeware (Free)
- WindowsAll
- The BGSSJ allows for easy and interactive querying using different gene identifiers (GenBank ID, UniGene, SwissProt, gene symbol), generates a summary page with listings of the frequencies of Gene Ontology annotations for each functional category. ...
- BGSSJ-Client-1.09a.jar
- bgssj
- Freeware (Free)
- 1.04 Mb
- N/A
- Magical Marker 1.7 is a stylistic, modern and powerful editors tool. Your monitor turns into a white board. Magical Marker draw something directly on the screen working with any application. You can do beautiful presentation using stamps and stickers. ...
- MagicalMarker US.sit
- MAKI Enterprise
- Commercial ($49.00)
- 819 Kb
- Mac OS Classic
- GMDR was built as an Open-Source interaction analysis instrument that is aimed to perform gene-gene interaction with generalized multifactor dimensionality methods.This Java-based tool offers a generalized combinatorial approach for detecting. ...
- gmdr.jar
- Guo-Bo Chen
- Freeware (Free)
- Windows NT, 2K, XP, Vista, 7
- Folder Marker is one of those little utilities that once you have it you wonder how you ever did with out it. It lets you mark your Windows Explorer folders in a couple of different ways, either using the context menu (right click) or by starting the. ...
- FolderMarkerFree.exe
- ArcticLine Software
- Freeware (Free)
- 983 Kb
- Win95, Win98, WinME, WinNT 4.x, WinXP, Windows2000, Windows2003
- The Gene Ontology Annotative Listing (GOAL) is an open-source PHP application for assembling and visualizing Biological Sequences based on their corresponding hierarchal Gene Ontology structure, described by the gene ontology. ...
- GOAL, Gene OntologyAnnotative Listing
- goal
- Freeware (Free)
- 14 Kb
- Linux
- The SPELL gene expression explorer searches multiple datasets for related gene expression patterns.Hibbs MA, et al. Exploring the functional landscape of gene expression: Directed search of large microarray compendia. Bioinformatics,. ...
- SPELL-src_2.0.2.tgz
- gene-spell
- Freeware (Free)
- 1.45 Mb
- N/A
- As you may guess from the name, Dart Marker is scorekeeping software for darts. The current version is called 'Dart Marker 3 - Cricket Scoreboard', and only scores for Cricket. The original Dart Marker Software was designed for DOS, and although. ...
- cricket.zip
- www.johnclare.com
- Freeware (Free)
- 67 Kb
- Windows XP, 2000, 98, Me, NT
- Gene Network Inference Analisys and Simulation. Infer gene relations from: 1) Literature: PubMed 2) Web: by quering search engines 3) Gene Expression Omnibus at NIH Then try to perform an analysis of the network dynamics.
- GeNIASim
- Nick V.
- Freeware (Free)
- Windows
- PRADA is designed for paired-end RNA sequencing data. PRADA focuses on the processing and analysis of mRNA profiles such as gene expression estimation and gene fusions identification. It implements several steps of analyses which are branched into two ...
- PRADA.zip
- Rahulsimham Vegesna, RoelVerhaak, Siyuan, WandalizTorres-Garcia
- Freeware (Free)
- 9.03 Mb
- Windows
Gene Marker Software For Mac Download
Related:Gene Marker Software For Mac Windows 10
Gene Marker - Gene Marker Download - Gene Marker Program - Marker Gene Technologies - Gene Marker MacGene Marker Software For Mac Free
Pages : <1 | 2 | 3
- Blog
- Home
- Best Free Accounting Software For Mac 2017
- Best Mac Document Scanner Software
- Theatre Script Writing Software Mac
- Where Is The Keychain App On A Mac
- Does Wd My Passport Software Work On Mac
- Roland Gx-24 Cutstudio Software Download Mac
- Best Internet Radio Software Mac
- Dji Inspire App For Mac
- Best 3d Print Software Mac
- Tv App Mac Os X
- Gene Marker Software For Mac
- Samsung S8 Unlock Mac Software
- Canon Mg6400 Scanner Software For Mac
- Remote For Mac Mini App
- Electronic Schematic Software Free Mac
- Add Calendat To Mac Today App
- Installing Software Mac Across All Accoumts
- Mac Os X Chromcast App 10.7.5
- Remove The Dymo App On My Mac
- Viper Mouse V560 Software Download Mac
- Free App To Transfer Files From Note 8 To Mac
- Mac App Check Disk Space
- Online Business Tax Software For Mac
- Comic Drawing Software Free Mac
- Mac Os 3d Printing Software
- Adobe After Effects Render Engine App Mac
- House Floor Plan App For Mac
- Tata Photon Software For Mac
- Spectrum App For Mac Pro
- Digital Mixer Software For Mac
- Free Relational Database Software For Mac
- Slideshow Software For Mac 2014
- Why Does Mac Book App Ruin The Book Format
- Free Smart Notebook Software For Mac
- Mac Place App On Desktop
- Photo Manipulation Software For Mac Free
- Texting Apps For Droid And Mac
- Electrical Schematic Software Free Mac
- When Apps Are Downloaded On Macs Where Do They Go
- Force Empty Trash Mac App
- Smart Board Software Download Mac
- Wps Office Presentation Software Mac
- Best Online 2d Animation Software For Mac
